Comparing DNA Sequences to Understand Evolutionary Relationships With Blast
How can bioinformatics be used as a tool to determine evolutionary relationships and to better understand genetic diseases?
In this laboratory investigation, you will use BLAST to compare several genes, and then use the information to construct a cladogram. A cladogram is treelike, with the endpoints of each branch representing a specific species. The closer two species are located to each other, the more recently they share a common ancestor. In the cladogram below, you will also see the shared derived characters. Note that the placement of the shared derived character corresponds to when (in a general, not a specific sense) that character evolved; every species above the character label possesses that structure.
Websites to use throughout lab:
NCBI- Gene site:
http://www.ncbi.nlm.nih.gov/gene
BLAST website:
http://blast.ncbi.nlm.nih.gov/Blast.cgi
Optional Online Activities to help you with this lab:
- “The Evolution of Flight in Birds” http://www.ucmp.berkeley.edu/education/explorations/reslab/flight/main.htm This activity provides a real-world example of how cladograms are used to understand evolutionary relationships.
- “What did T. rex taste like?” http://www.ucmp.berkeley.edu/education/explorations/tours/Trex/index.html
- “Journey into Phylogenetic Systematics” http://www.ucmp.berkeley.edu/clad/clad4.html
Procedure
Complete the Prelab Questions: BLASTPrelab

Little is known about the fossil. It appears to be a new species. Upon careful examination of the fossil, small amounts of soft tissue have been discovered. Normally, soft tissue does not survive fossilization; however, rare situations of such preservation do occur. Scientists were able to extract DNA nucleotides from the tissue and use the information to sequence several genes. Your task is to use BLAST to analyze these genes and determine the most likely placement of the fossil species on Figure 4.A team of scientists has uncovered the fossil specimen in Figure 3 near Liaoning Province, China. Make some general observations about the morphology (physical structure) of the fossil, and then record your observations in your notebook.
Step 1 Form an initial hypothesis as to where you believe the fossil specimen should be placed on the cladogram based on the morphological observations you made earlier. Draw your hypothesis on Figure 4.
Step 2 Locate and download gene files. Download three gene files from
http://blogging4biology.edublogs.org/2010/08/28/college-board-lab-files/.
Direct link to files:
Step 3 Upload the gene sequence into BLAST by doing the following:
a. Go to the BLAST homepage: http://blast.ncbi.nlm.nih.gov/Blast.cgi
b. Click on “Saved Strategies” from the menu at the top of the page.
Figure 5
c. Under “Upload Search Strategy,” click on “Browse” and locate one of the gene files you saved onto your computer.
d. Click “View.”
Figure 6
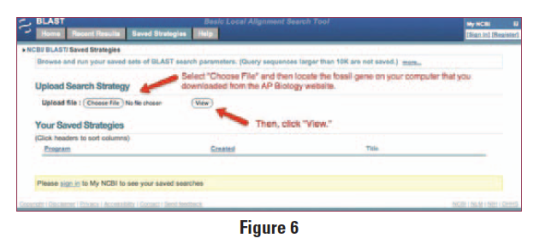 e. A screen will appear with the parameters for your query already configured. NOTE: Do not alter any of the parameters. Scroll down the page and click on the “BLAST” button at the bottom.
e. A screen will appear with the parameters for your query already configured. NOTE: Do not alter any of the parameters. Scroll down the page and click on the “BLAST” button at the bottom.
Figure 7
f. After collecting and analyzing all of the data for that particular gene (see instructions below), repeat this procedure for the other two gene sequences.
Step 4 The results page has two sections. The first section is a graphical display of the matching sequences.
Figure 8
Scroll down to the section titled “Sequences producing significant alignments.” The species in the list that appears below this section are those with sequences identical to or most similar to the gene of interest. The most similar sequences are listed first, and as you move down the list, the sequences become less similar to your gene of interest.
Figure 9
If you click on a particular species listed, you’ll get a full report that includes the classification scheme of the species, the research journal in which the gene was first reported, and the sequence of bases that appear to align with your gene of interest.
Figure 10
If you click on a particular species listed, you’ll get a full report that includes the species’ classification scheme, the research journal in which the gene was first reported, and the sequence of bases that appear to align with your gene of interest. If you click on the link titled “Distance tree of results,” you will see a cladogram with the species with similar sequences to your gene of interest placed on the cladogram according to how closely their matched gene aligns with your gene of interest.
Analyzing Results
Recall that species with common ancestry will share similar genes. The more similar genes two species have in common, the more recent their common ancestor and the closer the two species will be located on a cladogram.
As you collect information from BLAST for each of the gene files, you should be thinking about your original hypothesis and whether the data support or cause you to reject your original placement of the fossil species on the cladogram. For each BLAST query, consider the following:
- The higher the score, the closer the alignment.
- The lower the e value, the closer the alignment.
- Sequences with e values less than 1e-04 (1 x 10-4) can be considered related with an error rate of less than 0.01%.
- What species in the BLAST result has the most similar gene sequence to the gene of interest?
- Where is that species located on your cladogram?
- How similar is that gene sequence?
- What species has the next most similar gene sequence to the gene of interest?
- Based on what you have learned from the sequence analysis and what you know from the structure, decide where the new fossil species belongs on the cladogram with the other organisms. If necessary, redraw the cladogram you created before.
Evaluating Results
Compare and discuss your cladogram with your classmates. Does everyone agree with the placement of the fossil specimen? If not, what is the basis of the disagreement? On the main page of BLAST, click on the link “List All Genomic Databases.” How many genomes are currently available for making comparisons using BLAST? How does this limitation impact the proper analysis of the gene data used in this lab? What other data could be collected from the fossil specimen to help properly identify its evolutionary history?
Designing and Conducting Your Investigation
Now that you’ve completed this investigation, you should feel more comfortable using BLAST. The next step is to learn how to find and BLAST your own genes of interest. To locate a gene, you will go to the Entrez Gene website (http://www.ncbi.nlm.nih.gov/gene). Once you have found the gene on the website, you can copy the gene sequence and input it into a BLAST query.
Example Procedure
Your starting question may be: What is the function of actin in humans? Do other organisms have actin? If so, which ones?
- Go to the Entrez Gene website (http://www.ncbi.nlm.nih.gov/gene) and search for “human actin.”
- Click on the first link that appears and scroll down to the section “NCBI Reference Sequences.”
- Under “mRNA and Proteins,” click on the first file name. It will be named “NM001100.3” or something similar. These standardized numbers make cataloging sequence files easier. Do not worry about the file number for now.
- Just below the gene title click on “FASTA.” This is the name for a particular format for displaying sequences.
- The nucleotide sequence displayed is that of the actin gene in humans.
- Copy the entire gene sequence, and then go to the BLAST homepage (http://blast.ncbi.nlm.nih.gov/Blast.cgi).
- Click on “nucleotide blast” under the Basic BLAST menu.
- Paste the sequence into the box where it says “Enter Query Sequence.”
- Give the query a title in the box provided if you plan on saving it for later. S50 Investigation 3
- Under “Choose Search Set,” select whether you want to search the human genome only, mouse genome only, or all genomes available.
- Under “Program Selection,” choose whether or not you want highly similar sequences or somewhat similar sequences. Choosing somewhat similar sequences will provide you with more results.
- Click BLAST.
Below is a list of some gene suggestions you could investigate using BLAST. As you look at a particular gene, try to answer the following questions:
- What is the function in humans of the protein produced from that gene?
- Would you expect to find the same protein in other organisms? If so, which ones?
- Is it possible to find the same gene in two different kinds of organisms but not find the protein that is produced from that gene?
- If you found the same gene in all organisms you test, what does this suggest about the evolution of this gene in the history of life on earth?
- Does the use of DNA sequences in the study of evolutionary relationships mean that other characteristics are unimportant in such studies? Explain your answer.
Suggested Genes:
- ATP synthase
- Catalase
- GAPDH
- Keratin
- Myosin
- Pax1
- Ubiquitin







